Computational modeling uncovers new molecular mechanisms controlling T helper cell function
BLACKSBURG, Va., April 04, 2013 – Researchers at the Center for Modeling Immunity to Enteric Pathogens (MIEP) at Virginia Bioinformatics Institute (VBI) believe they have identified the mechanism that turns pro-inflammatory Th17 cells into anti-inflammatory T regulatory cells in the gut. This marks the first time that a comprehensive computational and mathematical model of the mechanisms governing Th cell differentiation has been created and key model-derived predictions have been validated experimentally. The results are published in PLOS Computational Biology.
The study focuses on PPAR γ, a protein that aids in metabolic regulation. In previous work, researchers at VBI’s Nutritional Immunology and Molecular Medicine Laboratory (NIMML) demonstrated that PPAR γ plays a crucial role in suppressing inflammation and immune responses. The new model developed by the MIEP team predicted a novel role of PPAR γ in controlling how gut-associated Th cells change from pro-inflammatory to anti-inflammatory types.
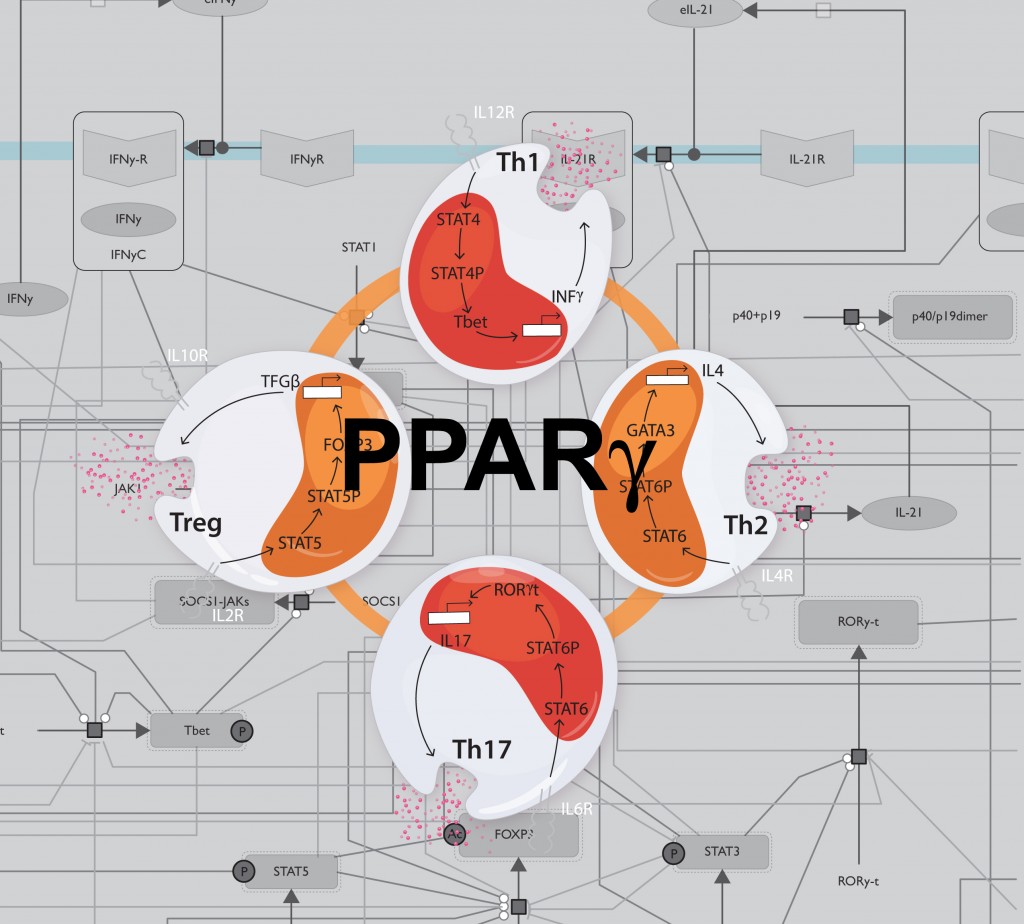 “Th cells play important roles in orchestrating immune responses, and their functions are controlled by complex signaling pathways with abundant feedback loops. In this project, we used computational and mathematical modeling to more comprehensively characterize the mechanisms controlling Th cell differentiation and plasticity at the systems level. We provide evidence demonstrating that PPAR γ activation controls the switch from pathogenic Th17 cells into regulatory T cells in the gut,” said Josep Bassaganya-Riera, professor of immunology, director of NIMML, and principal investigator of MIEP.
“Th cells play important roles in orchestrating immune responses, and their functions are controlled by complex signaling pathways with abundant feedback loops. In this project, we used computational and mathematical modeling to more comprehensively characterize the mechanisms controlling Th cell differentiation and plasticity at the systems level. We provide evidence demonstrating that PPAR γ activation controls the switch from pathogenic Th17 cells into regulatory T cells in the gut,” said Josep Bassaganya-Riera, professor of immunology, director of NIMML, and principal investigator of MIEP.
According to the researchers, the precision with which computational models unravel complex biological networks brings researchers closer to understanding how immune responses are shaped by genetics, environmental cues and disease states. The systems immunology approaches developed by MIEP may help researchers and clinicians identify targets that will yield more effective therapies for infectious and immune-mediated diseases.
Please visit the MIEP Web Portal at www.modelingimmunity.org.
MIEP is funded by the National Institute of Allergy and Infectious Diseases, part of the National Institutes of Health, under Contract No. HHSN272201000056C. PI: Josep Bassaganya-Riera.
About NIMML
The NIMML Institute is a 501 (c) (3) non-profit public charity foundation focused on a transdisciplinary, team-science approach to precision medicine at the interface of immunology, inflammation, and metabolism. The NIMML Institute team has led numerous large-scale transdisciplinary projects and is dedicated to solving important societal problems by combining the expertise of immunologists, computational biologists, toxicologists, modelers, translational researchers, and molecular biologists. The Institute is headquartered in Blacksburg, VA. For more information, please visit www.nimml.org or contact pio@nimml.org.
