New Computational model of host responses to Clostridium difficile infection
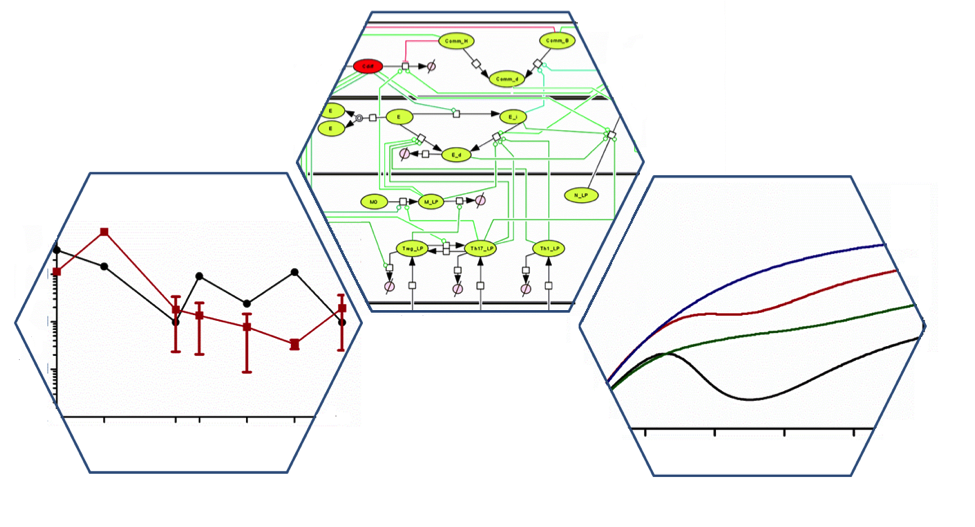
The MIEP team has published a computational model of interactions between C. difficile, the microbiome and the host. The article, entitled “Systems Modeling of Interactions between Mucosal Immunity and the Gut Microbiome during Clostridium difficile Infection”, was published in PLOS One on July 31.
The model follows the time course of infection moving from initiation to resolution based on immunological data acquired from a mouse model of C. difficile infection. Computational simulations illustrated the effect of antimicrobial peptides generated from epithelial cells and neutrophils on the clearance of C. difficile and also the regrowth of beneficial commensal bacteria following antibiotic reduction. The impaired regrowth of the colonic host microbiome by these antimicrobial peptides may help explain why high rates of recurrence occur after initial infections with C. difficile.
Additional analysis of the model demonstrated a crucial role for the balance between CD4+ T helper cell subsets, Th17 and iTreg, and neutrophilic influx on the development of damage to the epithelium, a key pathological outcome of disease. The innovative computational immunology approaches used to simulate the effects of the host microbiome may be used to explore similar mechanisms to other gastroenteric pathogens such as H. pylori and immune-mediated diseases such as Crohn’s disease or Ulcerative colitis. The computational model used in this study is available for download at Biomodels.net (MODEL1507200000).
About NIMML
The NIMML Institute is a 501 (c) (3) non-profit public charity foundation focused on a transdisciplinary, team-science approach to precision medicine at the interface of immunology, inflammation, and metabolism. The NIMML Institute team has led numerous large-scale transdisciplinary projects and is dedicated to solving important societal problems by combining the expertise of immunologists, computational biologists, toxicologists, modelers, translational researchers, and molecular biologists. The Institute is headquartered in Blacksburg, VA. For more information, please visit www.nimml.org or contact pio@nimml.org.
